染色质免疫沉淀(ChIP)在技术上具有挑战性,并且结果难以解读。Active Motif的ChIP IT®Express试剂盒通过应用蛋白质G偶联磁珠的简化方案来消除其中的一些挑战,使ChIP只需一天就能完成。试剂盒提供超声或者酶消化两种方案制备染色质。使用Active Motif提供的ChIP-IT®对照试剂盒和qPCR引物组引物组和已剪切好的染色质来完成您的实验设计,可以在每一步验证您的结果。要了解这些功能与我们提供的其他ChIP试剂盒的差异,请参阅我们的ChIP试剂盒选择指南。
ChIP-IT® Express试剂盒亮点
- ChIP-IT Express超声法对于难以裂解的细胞、培养细胞和组织样品来说是高质量的染色质片段化方案。
- 如无超声仪,培养细胞系样本只需简单的使用我们的ChIP-IT Express Enzymatic kit。
- 快速的操作步骤使ChIP实验在一天内完成成为可能。
- 蛋白质G磁珠提供低背景富集和简化的洗涤步骤。
要了解更多有关ChIP-IT Express的信息,请单击下面的“说明”和“概述选项卡。找专家做实验?我们的表观遗传服务团队可以为您提供ChIP、ChIP-Seq和许多其他基因组范围的数据生成和分析服务。
| Name | Format | Cat No. | Price | |
|---|---|---|---|---|
| ChIP-IT® Express | 25 rxns | 53008 | ¥7,800 | Add to Cart |
| ChIP-IT® Express Enzymatic | 25 rxns | 53009 | ¥8,260 | Add to Cart |
| ChIP-IT® Express Shearing Kit (included in 53008) | 10 rxns | 53032 | ¥3,710 | Add to Cart |
| ChIP-IT® Express Enzymatic Shearing Kit (included in 53009) | 10 rxns | 53035 | ¥4,230 | Add to Cart |
| ChIP-IT® Protein G Magnetic Beads (included in 53008 & 53009) | 25 rxns | 53014 | ¥4,160 | Add to Cart |
| Siliconized Tubes, 1.7 ml | 25 tubes | 53036 | ¥1,620 | Add to Cart |
ChIP-IT Express改善ChIP
ChIP-IT Express通过减少或消除几个耗时的步骤来改进传统ChIP。由于所提供的磁珠的背景比传统琼脂糖珠低得多,因此不再需要预先清理和封闭步骤。因为离心步骤已被快速磁性吸附取代,所以洗涤更容易。而且,与传统ChIP方案的8次洗涤相比,只需要3次洗涤。这大大减少了实验过程中的动手操作时间。这不仅节省了时间,而且减少了样本的操作次数,最大限度地减少了洗涤过程中可能发生的DNA损失,这有助于提高产量以及保持样本间的一致性。
优化的缓冲液减少背景
ChIP-IT Express试剂盒中包含的磁珠和磁条使ChIP更快和更容易,试剂盒的缓冲液也经过优化,为您提供更好的ChIP结果。洗脱和解交联缓冲液有助于减少背景,这使您能够识别较小的信号峰。背景的减少也使少量起始细胞的实验成为可能(请参见概览选项卡中的图2)。此外,这些缓冲液是规划一起使用的,所以您现在可以在从磁珠上洗脱DNA的同时进行解交联。
这些简单的改进使您能够快速、轻松地得到结果,让同时进行多个ChIP实验成为可能。使用ChIP IT Express试剂盒,您可以使用多通道移液器在PCR管中进行多个ChIP(图1和图2)。这种方法实际上使我们开发出了ChIP-IT Express HT试剂盒。这是一种高通量ChIP试剂盒,可以在96孔板上同时处理多达96个样本。
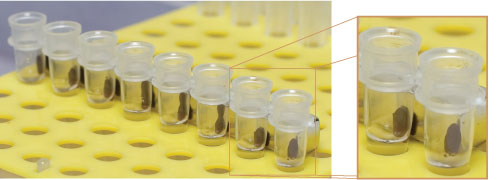

图1 & 2:使用ChIP-IT Express处理多个样本
磁珠的应用使得免疫沉淀和清洗步骤更加快速和容易,因为磁珠颗粒聚集在试管的一侧,甚至被完全拉出缓冲液。在不干扰磁珠颗粒的情况下去除缓冲液更简单,这使得现在使用多通道移液器在PCR管中执行多个ChIP成为可能。
强力磁条
ChIP-IT Express试剂盒包含一个强力条形磁铁,可将移液枪头盒转变为用于免疫沉淀和清洗的磁性支架。P1000枪头盒可与Eppendorf管一起使用(图3),而P200枪头盒可与PCR管一起使用(图1和2)。其他商用的磁性管支架,例如Ambion(图4)和Promega的支架也可以使用。
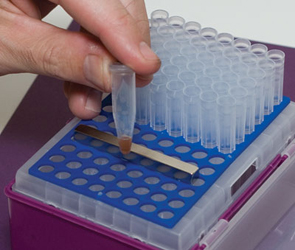
图3:使用标准枪头盒进行磁珠ChIP
ChIP-IT Express试剂盒包含一个强力条形磁铁,可将移液枪头盒转换为磁性支架,用于洗涤蛋白质G偶联的磁珠。

图4:ChIP-IT Express配套使用Ambion 6孔管磁性支架
商用的磁性管架,如Ambion和Promega的,也可以与ChIP-IT Express一起使用。
超声或可重复酶切方案的选择
成功的ChIP实验的第一步是将染色质打断成200-1000bp的片段。最常见的片段化方案是使用超声设备,如Active Motif的EpiShear™超声仪。然而,由于超声过程中乳化和过热的问题,使得超声方案很难优化。正因为如此,或者如果你没有超声仪,Active Motif已经开发出一种高效、温和的通过酶消化来剪切染色质的方案。因为Active Motif提供ChIP-IT-Express和ChIP-IT-Express Enzymatic试剂盒,所以您可以选择您喜欢的方法。更多完整信息,请点击此处了解。
ChIP配套产品和ChIP & ChIP-Seq验证抗体
为了简化实验问题排查、实验结果解读和验证抗体在ChIP中的可用性,Active Motif提供了物种特异性(人、小鼠和大鼠)ChIP-IT对照qPCR试剂盒,其中包含一个阳性对照抗体,一个增强小鼠单克隆抗体结合亲和力的桥连抗体,一个用于评估非特异性结合的阴性对照抗体,以及用于实时或终点PCR的物种特异性阳性和阴性对照引物。此外,我们还为人类、小鼠、大鼠、斑马鱼、果蝇和酵母中的许多常见ChIP靶点提供超过35种ChIP对照qPCR引物组。最后,Active Motif提供不断增加的ChIP验证抗体和ChIP-Seq验证抗体,这些抗体非常适合与我们所有的ChIP-IT试剂盒和ChIP相关产品一起使用。
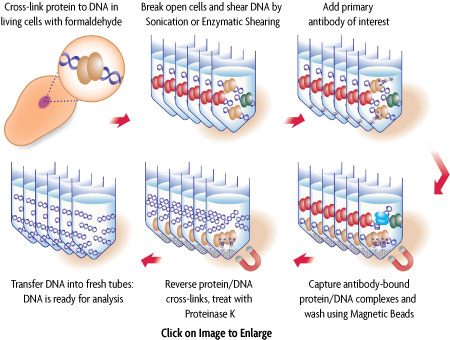
试剂盒是如何工作的?
ChIP涉及免疫沉淀经过交联固定的蛋白/DNA复合物。首先,完整的细胞用甲醛固定交联,从而保持蛋白和DNA的相互作用。然后,DNA被剪切成均匀的小片段,使用目标DNA结合蛋白的抗体免疫沉淀DNA/蛋白复合物。免疫沉淀后,DNA经过洗涤,解交联,蛋白被蛋白酶K处理去除。洗脱后的DNA可以用Active Motif的染色体免疫沉淀DNA纯化试剂盒进行纯化,纯化后的DNA可以直接进行qPCR或NGS测序下游分析。
图1:使用ChIP-IT Express的染色质免疫沉淀示意图
ChIP-IT Express提供高质量的ChIP-Seq数据
使用3µg Active Motif的AbFlex®组蛋白H3K9ac重组抗体(目录号91103)和Active Motif的ChIP-IT Express试剂盒免疫沉淀25µg来自K-562细胞的染色质。在ChIP之后,使用下一代DNA文库建库试剂盒(目录号53216和53264)制备Illumina兼容测序文库,并使用NextSeq 500进行测序。结果显示高质量的ChIP-Seq信号峰横跨1号染色体的一个区域。
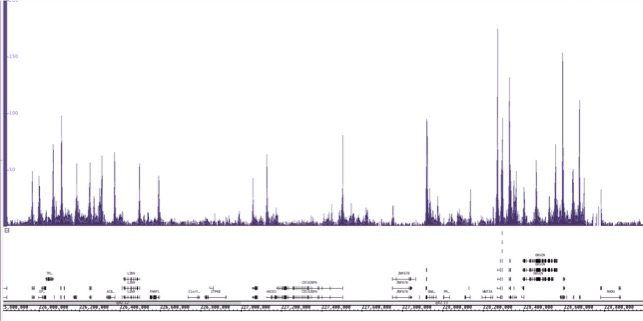
图2:使用ChIP-IT Express试剂盒免疫沉淀组蛋白修饰H3K9ac的ChIP-Seq数据
使用ChIP-IT Express富集组蛋白修饰H3K9ac的ChIP qPCR数据
使用3µg Active Motif的AbFlex®组蛋白修饰H3K9ac重组抗体(货号91103)和Active Motif的ChIP-IT Express试剂盒免疫沉淀25µg K-562细胞染色质。ChIP后,DNA按1:20稀释,使用Active Motif的人阳性对照引物组ACTB-1和GAPDH-1(货号号71003和71004),人阴性对照引物组1(货号71001)进行qPCR。结果以每1000个细胞检测到的结合情况展示,结果显示阳性对照PCR引物有富集,而阴性对照引物几乎没有富集。
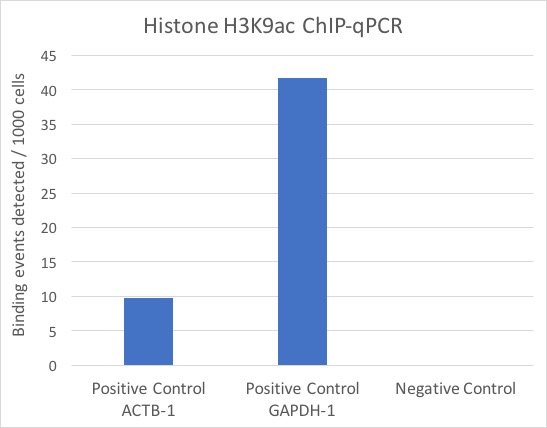
图3:使用ChIP-IT Express试剂盒免疫沉淀组蛋白修饰H3K9ac的ChIP-qPCR数据
灵活订货选项
ChIP-IT Express和ChIP-IT Express Enzymatic 试剂盒分别包括蛋白质G磁珠,ChIP-IT Shearing或Enzymatic Shearing试剂盒中的剪切试剂。然而,根据每个剪切染色质样本的ChIP反应数量,您可能会在用完ChIP-IT Express试剂盒中的所有免疫沉淀试剂之前用完剪切试剂。为此,ChIP-IT Sonication & Enzymatic Shearing试剂盒和蛋白质G磁珠也可以单独订购。
ChIP-IT Express试剂盒包含的试剂可进行25个ChIP反应。剪切试剂足够用于优化剪切条件及从三个15 cm的培养板(4.5 x 107个细胞)中进行5次染色质剪切制备;每次制备的染色质足够进行大约6次ChIP反应。我们建议您也购买适合您的样本类型的ChIP-IT对照试剂盒,因为使用经过验证的对照组有助于排查实验问题,解释您的结果并验证ChIP中使用的抗体。
组分 & 储存
蛋白质G偶联磁珠、条形磁铁、便签粘点、剪切缓冲液(或消化缓冲液和酶切混合物)、ChIP缓冲液1和2、洗脱缓冲液、解交联缓冲液、洗涤液和10X PCR缓冲液、10X PBS、10X 甘氨酸、1X 裂解缓冲液、蛋白酶抑制剂、PMSF、EDTA、蛋白酶K、蛋白酶K反应终止溶液和RNase A。试剂储存条件从室温到-20°C不等。了解完整的试剂组分及储存条件,请下载完整的试剂盒手册。所有试剂在正确温度下储存,保证6个月内稳定。
下面列出了使用我们的ChIP-IT®Express染色质免疫沉淀试剂盒的一些发表文章。
ChIP-IT® Express (Catalog #53008)
- “Sulforaphane reactivates cellular antioxidant defense by inducing Nrf2/ARE/Prdx6 activity during aging and oxidative stress.” by Kubo et al. (2017) Scientific Reports 7(14130): 1-17.
- “cGAS drives noncanonical-inflammasome activation in age-related macular degeneration.” by Kerur et al. (2018) Nature Medicine 24: 50-61.
- “An integrated transcriptome and epigenome analysis identifies a novel candidate gene for pancreatic cancer.” by Jia et al. (2013) BMC Med Genomics 6(33).
- “CXCL12 protects pancreatic β-cells from oxidative stress by a Nrf2-induced increase in catalase expression and activity.” by Dinić et al. (2016) Proc Jpn Acad Ser B Phys Biol Sci. 92(9): 436-454.
- “Novel computational analysis of protein binding array data identifies direct targets of Nkx2.2 in the pancreas.” by Hill et al. (2011) BMC Bioinformatics 12(62).
- “PARP-1 and YY1 are important novel regulators of CXCL12 gene transcription in rat pancreatic beta cells.” by Marković et al. (2013) PLoS ONE 8(3): e59679.
- “Synergistic activations of REG I α and REG I β promoters by IL-6 and Glucocorticoids through JAK/STAT pathway in human pancreatic β cells.” by Yamauchi et al. (2015) J Diabetes Res. Epub: 173058.
- “Localization of Double-Strand Break Repair Proteins to Viral Replication Compartments following Lytic Reactivation of Kaposi's Sarcoma-Associated Herpesvirus.” by Hollingworth et al. (2017) J Virol. 91(22): e00930-17.
- “Crosstalk between histone modifications indicates that inhibition of arginine methyltransferase CARM1 activity reverses HIV latency.” by Zhang et al. (2017) Nucleic Acids Res 45(16): 9348-9360.
- “The Replicative Consequences of Papillomavirus E2 Protein Binding to the Origin Replication Factor ORC2.” by DeSmet et al. (2016) PLoS Pathogens 12(10): e1005934.
- “Distinctive patterns of epigenetic marks are associated with promoter regions of mouse LINE-1 and LTR retrotransposons.” by Rangasamy. (2013) Mob DNA 4(1):27.
- “Phosphorylation State of ZFP24 Controls Oligodendrocyte Differentiation.” by Elbaz et al. (2018) Cell Rep 23(8):2254-2263.
- “Changes in chromatin state reveal ARNT2 at a node of a tumorigenic transcription factor signature driving glioblastoma cell aggressiveness.” by Bogeas et al. (2018) Acta Neuropathol 135(2):267-283.
- “Polymorphism in Tmem132d regulates expression and anxiety-related behavior through binding of RNA polymerase II complex.” by Naik et al. (2018) Transl Psychiatry 8(1):1.
- “MELK is a novel therapeutic target in high-risk neuroblastoma.” by Guan et al. (2018) Oncotarget 9(2): 2591–2602.
- “The embryonic type of SPP1 transcriptional regulation is re-activated in glioblastoma.” by Kijewska et al. (2018) Oncotarget 8(10):16340-16355.
- “BIX01294, an inhibitor of histone methyltransferase, induces autophagy-dependent differentiation of glioma stem-like cells.” by Ciechomska et al. (2016) Scientific Reports 6(38723).
- “Enhancing dopaminergic signaling and histone acetylation promotes long-term rescue of deficient fear extinction.” by Whittle et al. (2016) Transl Psychiatry 6(12):e974.
- “Tissue mechanics promote IDH1-dependent HIF1α-tenascin C feedback to regulate glioblastoma aggression.” by Miroshnikova et al. (2016) Nat Cell Bio 18(12):1336-1345.
ChIP-IT® Express Enzymatic (Catalog #53009)
- “Polymorphism rs7278468 is associated with Age-related cataract through decreasing transcriptional activity of the CRYAA promoter.” by Ma et al. (2016) Scientific Reports 6(23206).
- “Deep sequencing and in silico analyses identify MYB-regulated gene networks and signaling pathways in pancreatic cancer.” by Azim et al. (2016) Sci. Rep. 6(28446).
- “Overexpression of IFN-induced protein with tetratricopeptide repeats 3 (IFIT3) in pancreatic cancer: cellular "pseudoinflammation" contributing to an aggressive phenotype.” by Niess et al. (2015) Oncotarget 6(5): 3306-3318.
- “Peretinoin, an Acyclic Retinoid, Inhibits Hepatitis B Virus Replication by Suppressing Sphingosine Metabolic Pathway In Vitro.” by Murai et al. (2018) Int J Mol Sci. 19(2): E108.
- “LIMD1 is induced by and required for LMP1 signaling, and protects EBV-transformed cells from DNA damage-induced cell death.” by Wang et al. (2018) Oncotarget 9(5): 6282–6297.
- “Human Beta Defensin 2 Selectively Inhibits HIV-1 in Highly Permissive CCR6⁺CD4⁺ T Cells.” by Lafferty et al. (2017) Viruses 9(5): E111.
- “CCCTC-binding factor recruitment to the early region of the human papillomavirus 18 genome regulates viral oncogene expression.” by Paris et al. (2015) J Virol. 89(9):4770-85.
- “Expansion of a novel endogenous retrovirus throughout the pericentromeres of modern humans.” by Zahn et al. (2015) Genome Biol 16:74.
- “HIV-1 induced nuclear factor I-B (NF-IB) expression negatively regulates HIV-1 replication through interaction with the long terminal repeat region.” by Vemula et al. (2015) Viruses 7(2):543-58.
- “Inflammation-induced miRNA-155 inhibits self-renewal of neural stem cells via suppression of CCAAT/enhancer binding protein β (C/EBPβ) expression.” by Obora et al. (2017) Sci Rep 7:43604.
- “BH3 mimetics suppress CXCL12 expression in human malignant peripheral nerve sheath tumor cells.” by Graham et al. (2017) Oncotarget 8(5): 8670–8678.
ChIP-IT® Protein G Magnetic Beads (included in Catalog #53008 & #53009)
- “RNA polymerase II primes Polycomb-repressed developmental genes throughout terminal neuronal differentiation.” by Ferrai et al. (2017) Mol Syst Biol 13(10):946.