Low Cell ChIP-Seq Overview
This product has been discontinued.
The Low Cell ChIP Kit is now offered without the NGS Library Preparation Kit and NGS Indexing Kit, enabling you to choose the NGS Kits best for your needs.
Active Motif's Low Cell ChIP-Seq Kit provides a complete ChIP-Seq workflow for chromatin preparation, immunoprecipitation, and next-generation sequencing (NGS) library preparation from limited amounts of cells or small tissue biopsies. To see how these features compare to the other ChIP kits we offer, see our ChIP Kit Selection Guide.
The assay is robust enough to work with both high abundance histone modifications and low abundance transcription factor proteins. The Next Gen DNA Library and Next Gen DNA Indexing reagents are included to generate high complexity sequencing libraries that are compatible with Illumina® platforms.
ChIP-Seq is a method that has traditionally required using millions of cells per immunoprecipitation reaction to generate genome-wide profiles for histone modifications or transcription factor binding. The Low Cell ChIP-Seq Kit not only reduces sample input requirements, but it also resolves issues often associated with low cell ChIP, including poor signal-to-noise, inefficient library amplification, and high duplication rates.
Low Cell ChIP-Seq Highlights:
- Reproducible ChIP-Seq data for histone modifications or transcription factors from as few as 1,000 cells
- Complete ChIP-Seq workflow includes chromatin preparation, immunoprecipitation, and Illumina-compatible NGS library preparation reagents that have been optimized for use on limited cell numbers
- Blocking reagents and an optimized protocol improve signal-to-noise ratio for better peak calling and lower background
- Molecular identifiers (MIDs) are incorporated within the P5 adapter for retention of fragmentation duplicates and removal of PCR duplicates to increase the number of unique alignments
- Multiplex up to 16 samples on the same sequencing flow cell
The Low Cell ChIP-Seq Kit contains enough reagents to perform 16 ChIP-seq reactions. Some of the included ChIP buffers, blockers, and protein G capture reagents may also be purchased separately.
First Time Using the Low Cell ChIP-Seq Kit?
Ensure your success with the Low Cell Optimization Module!
The Low Cell ChIP-Seq Optimization Module allows researchers to monitor the success of each critical step in the Low Cell ChIP-Seq protocol and optimize the experiments before using their valuable samples. This optimization module also enables scientists to evaluate the quality of chromatin preparations from low cell numbers to ensure best results in ChIP-Seq assays with low cell numbers.
Low Cell ChIP-Seq Contents & Storage
Please note that the Low Cell ChIP-Seq Kits are shipped on dry ice and contains reagents with multiple storage temperatures inside. All components can be stored at -20°C prior to first use, then we recommend storing each component at the temperatures indicated below. All reagents are guaranteed stable for 6 months from date of receipt when stored properly. Each kit includes the following components:
Reagents for Low Cell ChIP-Seq Kit
- 10X PBS; Store at -20°C
- Proteinase K (10 µg/µl); Store at -20°C
- Blocker; Store at -20°C
- Blocking Reagent AM1; Store at -20°C
- BSA; Store at -20°C
- 100 mM PMSF; Store at -20°C
- Protease Inhibitor Cocktail (PIC); Store at -20°C
- Carrier; Store at -20°C
- Fixation Buffer; Store at 4°C
- Protein G Agarose beads; Store at 4°C
- TE, pH 8.0; Store at RT
- 5 M NaCl; Store at RT
- Stop Solution; Store at RT
- ChIP Filtration Columns; Store at RT
- ChIP Buffer; Store at RT
- Wash Buffer AM1; Store at RT
- Elution Buffer AM4; Store at RT
- LiCl Buffer; Store at RT
- 1.7 ml siliconized tubes; Store at RT
Reagents for Next Gen DNA Library Kit* (included)
- Buffer W1; Store at -20°C
- Enzyme W2; Store at -20°C
- Buffer G1; Store at -20°C
- Reagent G2; Store at -20°C
- Enzyme G3; Store at -20°C
- Enzyme G4; Store at -20°C
- Buffer Y1; Store at -20°C
- Enzyme Y3 Store at -20°C
- Buffer B1; Store at -20°C
- Reagent B3; Store at -20°C
- Enzyme B4; Store at -20°C
- Enzyme B5; Store at -20°C
- Enzyme B6; Store at -20°C
- Reagent R2; Store at -20°C
- Buffer R3; Store at -20°C
- Enzyme R4; Store at -20°C
- Alu 115; Store at -20°C
- Alu 247; Store at -20°C
- PEG Solution; Store at RT
- Low EDTA TE; Store at RT
Reagents for Next Gen Indexing Kit* (included)
- Reagent Y2 Index 1; Store at -20°C
- Reagent Y2 Index 2; Store at -20°C
- Reagent Y2 Index 3; Store at -20°C
- Reagent Y2 Index 4; Store at -20°C
- Reagent Y2 Index 5; Store at -20°C
- Reagent Y2 Index 6; Store at -20°C
- Reagent Y2 Index 7; Store at -20°C
- Reagent Y2 Index 8; Store at -20°C
- Reagent Y2 Index 9; Store at -20°C
- Reagent Y2 Index 12; Store at -20°C
- Reagent Y2 Index 13; Store at -20°C
- Reagent Y2 Index 14; Store at -20°C
- Reagent Y2 Index 15; Store at -20°C
- Reagent Y2 Index 16; Store at -20°C
- Reagent Y2 Index 18; Store at -20°C
- Reagent Y2 Index 19; Store at -20°C
- Reagent R1; Store at -20°C
- Reagent B2 MID; Store at -20°C
* The Next Gen DNA Library and Next Gen Indexing Kits are powered by Swift Biosciences.
Low Cell ChIP-Seq Data
What is Low Cell Number?
With the Low Cell ChIP-Seq Kit, researchers can perform ChIP-Seq assays on samples with limited cell numbers that were previously incompatible with ChIP. The kit is ideal if working with limited sample amounts, hard to culture cell lines, or small tissue biopsies.
The recommended cell number listed below is the amount of cells that will be used for each chromatin preparation. The entire chromatin sample is then used in a single immunoprecipitation reaction. This differs from other ChIP assays where chromatin must be prepared from larger cell numbers and then the chromatin is diluted for use in the immunoprecipitation.
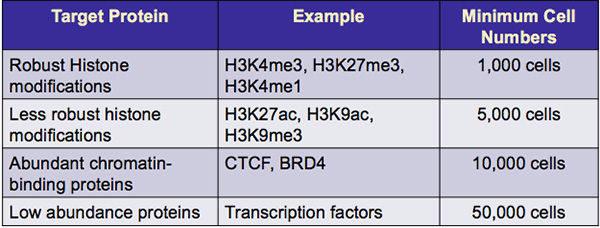
Table 1: Minimum recommended cell numbers for use with Low Cell ChIP-Seq Kit. Minimum cell numbers suggested are based on target protein abundance and antibody affinity.
Other factors that will influence the recommended cell number include cell type, abundance of target protein, and the quality of the ChIP-Seq antibody. Below are some validated antibodies from Active Motif for use with the Low Cell ChIP-Seq Kit.
| Protein Target | Active Motif Catalog Number | Cell Type Tested |
|---|---|---|
| H3K4me3 | 39159 | MDA-MB-468; GM12878 |
| H3K9ac | 39918 | GM12878 |
| H3K9me3 | 39161 | GM12878 |
| H3K27ac | 39133 | HepG2 |
| H3K27me3 | 39155 | MDA-MB-468 |
| CTCF | 61311 | MDA-MB-468 |
| BRD4 | 91301 | MDA-MB-468 |
Table 2: Active Motif's Low Cell ChIP-Seq validated antibodies.
Active Motif's Low Cell ChIP-Seq Kit was used to generate ChIP-Seq data from 10,000 MDA-MB-468 cells using an antibody directed against bromodomain-containing 4 (BRD4) protein, or 5,000 GM12878 cells using antibodies directed against histone H3K27ac or histone H3K27me3. Following ChIP, Illumina-compatible sequencing libraries were prepared using the included Next Gen DNA Library Kit and Next Gen Indexing Kits and sequenced using the NextSeq 500. Results show high quality ChIP-Seq peaks can be obtainined using as few as 5,000 cells for both active and repressive histone modifications and as few as 10,000 cells for BRD4. Peaks were compared to ENCODE data sets using 20 million cells.
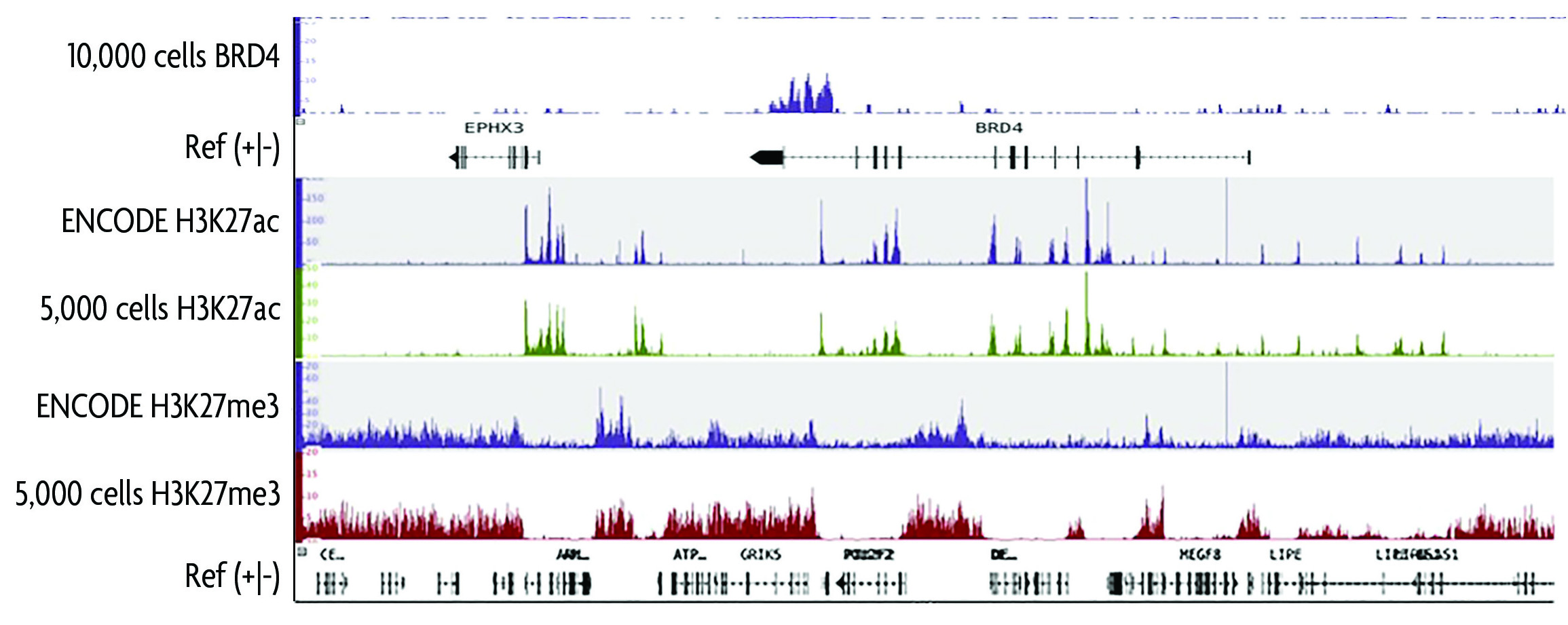
Figure 1: Low Cell ChIP-Seq peaks for BRD4, H3K27ac, and H3K27me3 compared to ENCODE data sets.
Below is a summary of Low Cell ChIP-Seq Kit data sets using both robust and low abundance target proteins. Nice peaks above background were observed from as few as 1,000 cells for the histone modifications H3K4me3, H3K4me1, and H3K27me3. Both active and repressive histone modifications are shown, indicating that the method is sensitive enough to obtain good ChIP-Seq signal even when working with repressive chromatin. Lower abundance proteins, including BRD4 and insulator protein CTCF showed nice ChIP enrichment from as few as 10,000 cells. Results shown are from a single Low Cell ChIP-Seq Kit reaction, however, data has been reproducibly generated across multiple experiments.
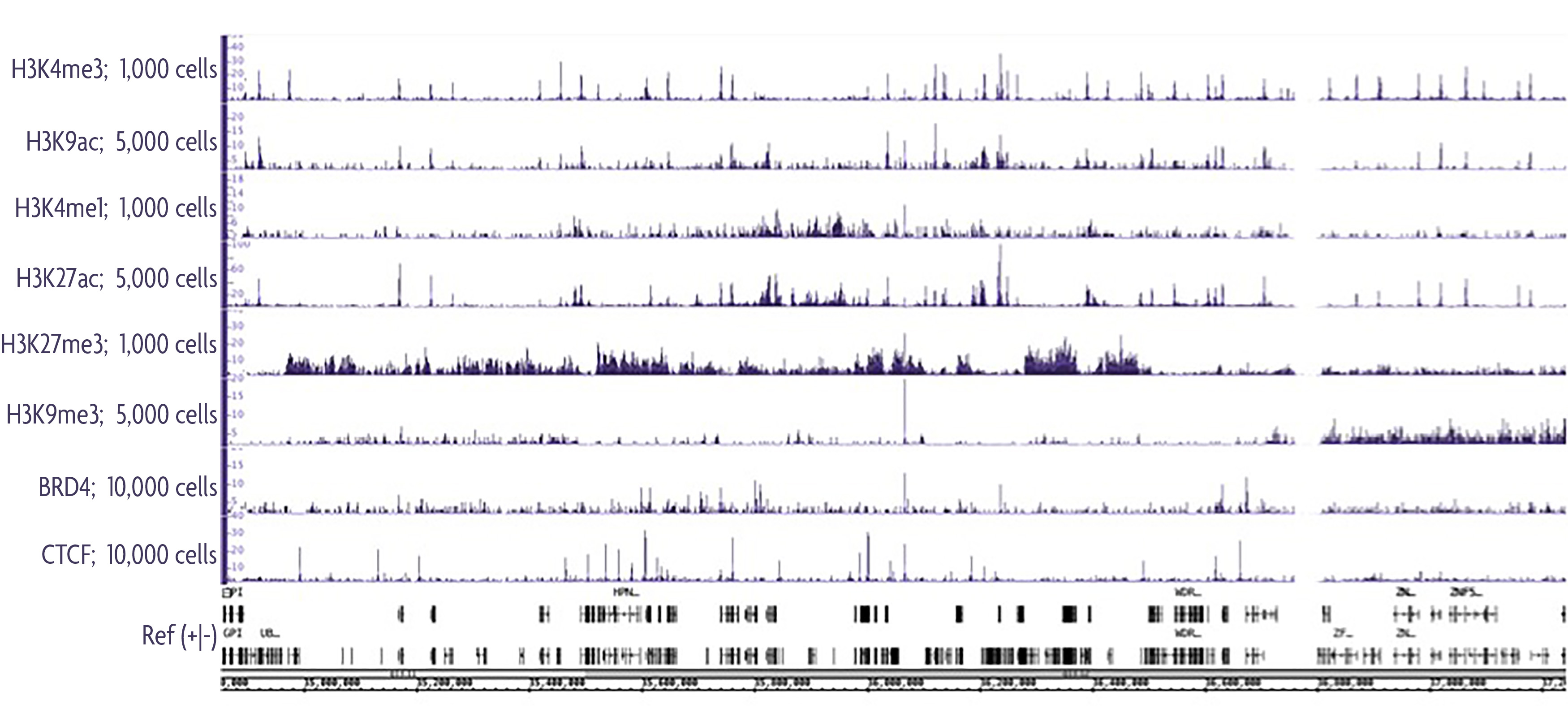
Figure 2: Summary of various Low Cell ChIP-Seq Kit experiments targeting both histone and low abundance proteins.
Low Cell ChIP-Seq Documents
You might also be interested in:
| Name | Format | Cat No. | Price | |
|---|---|---|---|---|
| Low Cell ChIP-Seq Kit | 16 rxns | 53084 | Discontinued | |
| ChIP Buffer | 50 ml | 37516 | ¥1,760 | Add to Cart |
| Blocking Reagent AM1 | 0.1 ml | 37496 | ¥1,760 | Add to Cart |
| BSA (10 mg/ml) | 0.1 ml | 37497 | ¥1,760 | Add to Cart |
| Blocker | 0.1 ml | 37498 | ¥1,760 | Add to Cart |
| Protein G Agarose Beads | 1.2 ml | 37499 | ¥2,670 | Add to Cart |
| TE, pH 8.0 | 35 ml | 37515 | ¥1,760 | Add to Cart |
What is Low Cell Number?
With Low Cell ChIP-Seq, researchers can reduce the number of cells they are studying to improve their understanding of the complexities of protein-DNA interactions to a more limited population of cells. The assay is ideal if working with limited sample amounts, hard to culture cell lines, or small tissue biopsies. The recommended cell number is the amount of cells that will be used for each chromatin preparation. The entire chromatin sample is then used in a single immunoprecipitation reaction. This differs from other ChIP assays where chromatin must be prepared from larger cell numbers and then the chromatin is diluted for use in the immunoprecipitation.

Figure 1: Table of the minimum recommended cell numbers for use with Low Cell ChIP-Seq Kit. Minimum cell numbers suggested are based on target protein abundance and antibody affinity.
Other factors that will influence the recommended cell number include cell type, abundance of target protein and the quality of the ChIP-Seq antibody. Below are some validated antibodies from Active Motif for use with Low Cell ChIP-Seq.
| Protein Target | Active Motif Catalog Number | Cell Type Tested |
|---|---|---|
| H3K4me3 | 39159 | MDA-MB-468; GM12878 |
| H3K9ac | 39918 | GM12878 |
| H3K9me3 | 39161 | GM12878 |
| H3K27ac | 39133 | HepG2 |
| H3K27me3 | 39155 | MDA-MB-468 |
| CTCF | 61311 | MDA-MB-468 |
Figure 2: Table of Active Motif's Low Cell ChIP-Seq validated antibodies.
Low Cell ChIP-Seq Data
Active Motif's Low Cell ChIP-Seq Kit was used to immunoprecipitate 10,000 MDA-MB-468 cells using an antibody directed against Bromodomain containing 4 (BRD4) protein, or 5,000 GM12878 cells using antibodies directed against Histone H3K27ac or Histone H3K27me3. Following ChIP, Illumina-compatible sequencing libraries were prepared using the included Next Gen DNA Library Kit and Next Gen Indexing Kits and sequenced using the NextSeq 500. Results show high quality ChIP-seq peaks can be obtainined using as little as 5,000 cells for both active and repressive histone modifications and as low as 10,000 cells for BRD4. Peaks were compared to ENCODE data sets using 20 million cells.

Figure 3: Low Cell ChIP-Seq peaks for BRD4, H3K27ac and H3K27me3 histone modifications compared to ENCODE data sets.
Below is a summary of Low Cell ChIP-Seq data sets using both robust and low abundance target proteins. Nice peaks above background were observed from as little as 1,000 cells for histone marks H3K4me3, H3K4me1 and H3K27me3. Both active and repressive histone modifications are shown, indicating that the method is sensitive enough to obtain good ChIP-Seq signal even when working with repressive chromatin. Lower abundance proteins, including BRD4 and insulator protein CTCF showed nice ChIP enrichment from as low as 10,000 cells. Results shown are from a single Low Cell ChIP-Seq reaction, however, data has been reproducibly generated across multiple experiments.

Figure 4: Summary of various Low Cell ChIP-Seq experiments targeting both histone and low abundance proteins.
The following publications cite the use of and/or provide additional information about the Low Cell ChIP-Seq Kit:
- Bcl11b is essential for licensing Th2 differentiation during helminth infection and allergic asthma. Lorentsen et al. (2018) Nature Communications
Contents & Storage
Please note that the Low Cell ChIP-Seq Kits are shipped on dry ice and contains reagents with multiple storage temperatures inside. All components can be stored at -20°C prior to first use, then we recommend storing each component at the temperatures indicated below. All reagents are guaranteed stable for 6 months from date of receipt when stored properly. Each kit includes the following components:
Reagents for Low Cell ChIP-Seq Kit
- 10X PBS; Store at -20°C
- Proteinase K (10 µg/µl); Store at -20°C
- Blocker; Store at -20°C
- Blocking Reagent AM1; Store at -20°C
- BSA; Store at -20°C
- 100 mM PMSF; Store at -20°C
- Protease Inhibitor Cocktail (PIC); Store at -20°C
- Carrier; Store at -20°C
- Fixation Buffer; Store at 4°C
- Protein G Agarose beads; Store at 4°C
- TE, pH 8.0; Store at RT
- 5 M NaCl; Store at RT
- Stop Solution; Store at RT
- ChIP Filtration Columns; Store at RT
- ChIP Buffer; Store at RT
- Wash Buffer AM1; Store at RT
- Elution Buffer AM4; Store at RT
- LiCl Buffer; Store at RT
- 1.7 ml siliconized tubes; Store at RT
Reagents for Next Gen DNA Library Kit* (included)
- Buffer W1; Store at -20°C
- Enzyme W2; Store at -20°C
- Buffer G1; Store at -20°C
- Reagent G2; Store at -20°C
- Enzyme G3; Store at -20°C
- Enzyme G4; Store at -20°C
- Buffer Y1; Store at -20°C
- Enzyme Y3 Store at -20°C
- Buffer B1; Store at -20°C
- Reagent B3; Store at -20°C
- Enzyme B4; Store at -20°C
- Enzyme B5; Store at -20°C
- Enzyme B6; Store at -20°C
- Reagent R2; Store at -20°C
- Buffer R3; Store at -20°C
- Enzyme R4; Store at -20°C
- Alu 115; Store at -20°C
- Alu 247; Store at -20°C
- PEG Solution; Store at RT
- Low EDTA TE; Store at RT
Reagents for Next Gen Indexing Kit* (included)
- Reagent Y2 Index 1; Store at -20°C
- Reagent Y2 Index 2; Store at -20°C
- Reagent Y2 Index 3; Store at -20°C
- Reagent Y2 Index 4; Store at -20°C
- Reagent Y2 Index 5; Store at -20°C
- Reagent Y2 Index 6; Store at -20°C
- Reagent Y2 Index 7; Store at -20°C
- Reagent Y2 Index 8; Store at -20°C
- Reagent Y2 Index 9; Store at -20°C
- Reagent Y2 Index 12; Store at -20°C
- Reagent Y2 Index 13; Store at -20°C
- Reagent Y2 Index 14; Store at -20°C
- Reagent Y2 Index 15; Store at -20°C
- Reagent Y2 Index 16; Store at -20°C
- Reagent Y2 Index 18; Store at -20°C
- Reagent Y2 Index 19; Store at -20°C
- Reagent R1; Store at -20°C
- Reagent B2 MID; Store at -20°C
* The Next Gen DNA Library and Next Gen Indexing Kits are powered by Swift Biosciences.


