
Single-Cell 10x Genomics Multiome Service Overview
Active Motif has been designated by 10x Genomics as a Certified Service Provider of Single-Cell Multiome. Single-Cell Multiome combines single-cell gene expression analysis with single-cell open chromatin mapping to provide genome-wide mapping of both the transcriptomal and epigenetic landscapes at the single-cell level. Understanding both the gene expression profile and the chromatin state at single-cell resolution can help identify how epigenetic changes instruct gene expression in distinct cell populations.

What are the advantages of using Single-Cell Multiome?
Single-Cell Multiome can be used to identify cell subpopulations with different transcriptome and epigenetic profiles within complex samples, eliminating the need for isolation strategies like FACS or magnetic sorting that could alter the biology due to sample manipulation. For example:
- Identifying novel cell that modulate response to drug treatments (i.e. responders vs. resistant cells)
- Identifying subpopulations of cells with variations in gene expression that can provide insight into developmental trajectories (i.e. brain development, T-helper cell development, B-cell differentiation)

Active Motif’s end-to-end Single-Cell 10x Genomics Multiome Service includes:
- Preparation of nuclei
- Tn5 tagmentation of open chromatin regions
- Nuclei loaded onto 10x Genomics Chromium Controller
- Barcoding of mRNA and in the same nucleus
- Library generation
- Sequencing
- Bioinformatic analysis
Join the Leading Researchers Using Active Motif's Single-Cell 10x Genomics Multiome Services














Single-Cell 10x Genomics Multiome Service Data
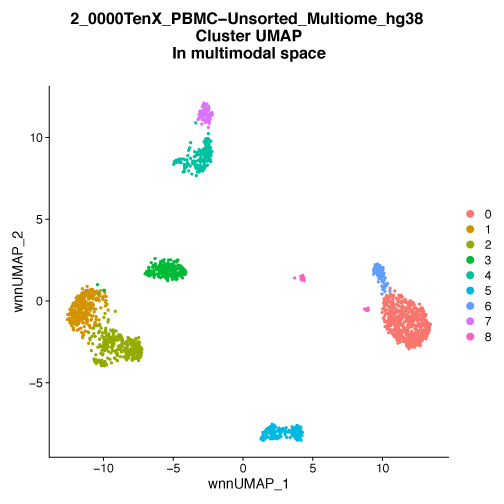
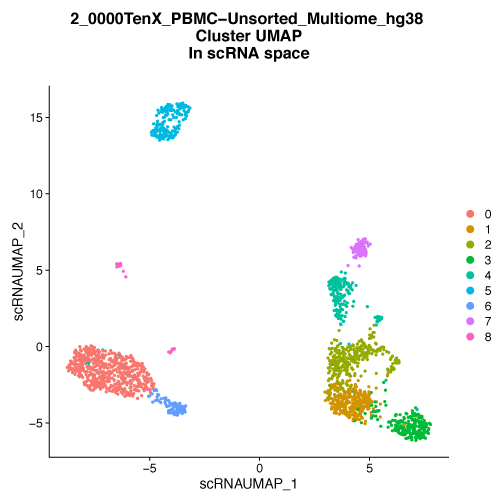
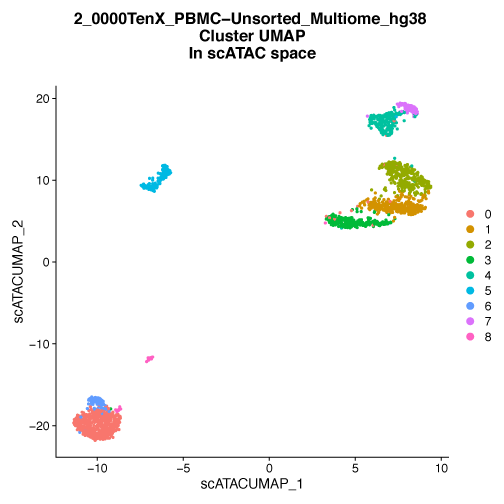
Figure 1. UMAP embedding of PBMC cells colored by multimodal cluster identity.
A) PBMC are plotted on a low dimensional embedding using the UMAP technique. Each dot is an individual cell and is plotted in two dimensions using the combination of both gene expression (scRNA-Seq) and chromatin accessibility (scATAC-Seq) data. The color of the dots indicates the cluster assignment of the cell, cells with the same color have similar patterns of gene expression and similar patterns of chromatin accessibility. These clusters are typically representative of cell types or cell states. Using both gene expression and chromatin accessibility data allows for the clear identification of well-separated clusters. B) UMAP embedding of the same cells as A, but in this case the embedding is based on gene expression data alone. The cluster assignments are identical to A and are based on both gene expression and chromatin accessibility data. When examining scRNA-Seq data in isolation the defined clusters lose some of the spatial separation seen in the multimodal analysis. C) UMAP embedding of the same cells as A based on chromatin accessibility alone. As with A and B, the cluster assignments are based on both gene expression and chromatin accessibility data. Examining cells by their chromatin accessibility alone also leads to a decrease in the separation of some clusters on this UMAP plot. Data provided by 10X Genomics.
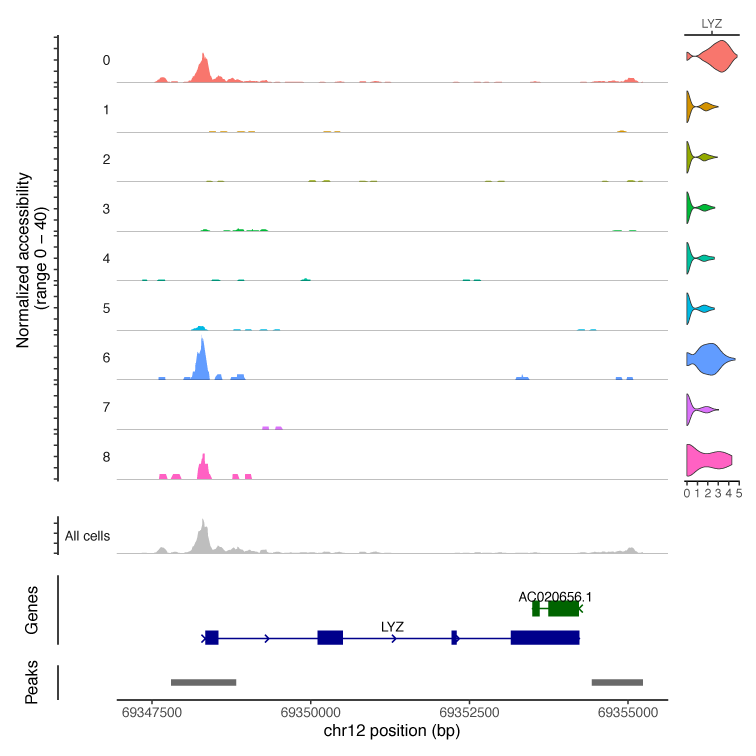
Figure 2. Expression and accessibility of the LYZ gene in PBMC clusters.
The bottom of the plot shows the LYZ gene body and the location of called scATAC-Seq peaks. The first 8 rows on the left show the chromatin accessibility profile of each cell cluster, and the ninth row shows a mock bulk ATAC-Seq profile. If these cells were examined using a traditional bulk ATAC-Seq method (gray track), a peak at the promoter would have been identified but it would not be possible to determine if that peak is found in all cells or specific cell types. The single cell analysis reveals that only cell clusters 0 (peach), 6 (blue), and 8 (peak) have open chromatin at the start of the LYZ gene. The violin plots on the right show the amount of LYZ expression in all cells for a given cluster. In agreement with the chromatin accessibility data, clusters 0, 6, and 8 have the highest LYZ expression. Data provided by 10X Genomics.
Single-Cell 10x Genomics Multiome Service FAQs
What is Single-Cell Multiome?
Single-Cell Multiome (scRNA+scATAC) measures both gene expression and maps open chromatin from same cell.
How is single-cell sequencing different from bulk sequencing?
Single-Cell Multiome measures gene expression and open chromatin at single cell resolution such that when examining a heterogenous cell population it can identify variations in gene expression between cell subpopulations within a single sample. Bulk RNA-Seq and ATAC-Seq measure the average expression level or chromatin open/closed state for each gene or locus across a large population of input cells.
Is this service specific to human?
Single-Cell Multiome is compatible with both human and mouse samples. Other species are bioinformatically unsupported (Cell Ranger software is limited to human and mouse).
Is Single-Cell Multiome compatible with FFPE samples?
No. This assay is not compatible with FFPE samples.
How many cells are required for Active Motif's Single-Cell Multiome service?
We ask for 250,000 to 2 million cryopreserved cells or 20-50 mg frozen tissue
What is the read depth for Active Motif's Single-Cell 10x Genomics Multiome Service?
Per 10X Genomics recommendations, we perform ~30,000 read pairs per cell, targeting 5000 cells.
Single-Cell 10x Genomics Multiome Service Documents
Single-Cell 10x Genomics Multiome Service Sample Submission Portal
Our online sample submission portal allows you to easily upload your service project samples and track your project status. Follow the sample submission instructions in the portal to ensure that all your samples arrive at Active Motif in the best possible condition and properly associated with your project.

