ATAC-Seq是基于转座酶介导,把引物插入开放的染色质区域的测序。 该实验提供了染色质开放和可及区域的全基因组图谱,这些区域指示了活跃的调节区域。
为什么要研究开放染色质?
- 深入了解基因调控,细胞对治疗或疾病的反应
- 识别哪些转录因子正在驱动细胞命运,疾病或反应
- 组织或原代细胞,例如胰岛β细胞
- 有限的病人样本
- 根据开放的染色质特征对患者或样品进行分层

当细胞数目受限时,如果尚不清楚表观遗传学是否在您的细胞系统的反应中起作用,或者不清楚使用ChIP-Seq研究哪种组蛋白修饰最重要,则ATAC-Seq可以很好地替代ChIP-Seq。
ATAC-Seq实验包括
- 细胞制备
- 转座酶反应
- 文库扩增
- 在Illumina平台上测序
- 生信分析
样本分型
Active Motif的服务团队是唯一一个经常从组织生成ATAC-Seq数据的小组。 Active Motif将提供以下样本类型的服务:
- 人和动物组织(包括异种移植物和人体活检组织)
- 原代细胞(包括T细胞和B细胞)
- 流式分选细胞
- 大多数罕见细胞群体
Sample Submission Portal
Our online sample submission portal allows you to easily upload your service project samples and track your project status. Follow the sample submission instructions in the portal to ensure that all your samples arrive at Active Motif in the best possible condition and properly associated with your project.
要了解更多信息,请向我们发送表观遗传服务信息申请。 您还可以下载Active Motif的表观遗传服务手册。
客户对我们的评价
"我正在研究心力衰竭的表观遗传学调控。我在Active Motif表观遗传学服务方面拥有非常好的体验,今后我将继续与Active Motif合作。我得到了销售部和技术支持团队的大力支持, 帮助我完成服务的所有项目。"
Ning Feng, MD, PhD
University of Pittsburgh
查看客户评价的完整列表 >
| Name | Cat No. | Price | |
|---|---|---|---|
| ATAC-Seq Service | 25079 | Get Quote | |
| Active Motif Epigenetic Services Brochure |
| Comprehensive ATAC-Seq Solutions Brochure |
| ATAC-Seq Sample Preparation |
| Epigenetic Services Citations |
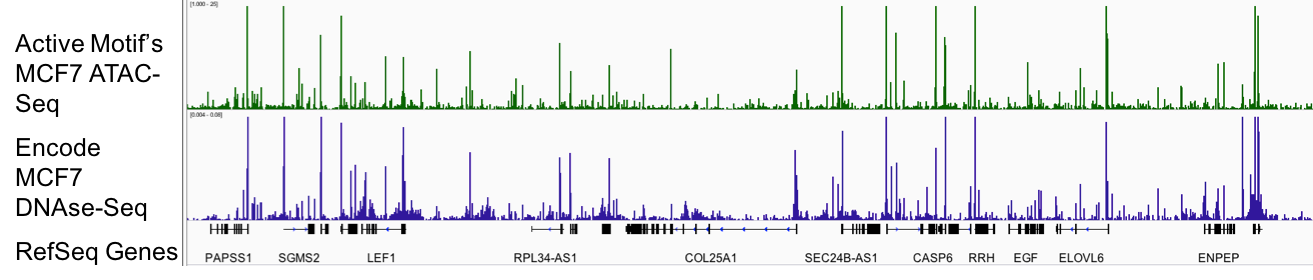
图1:Active Motif的ATAC-Seq分析能够可靠地检测出开放染色质区域。
DNAse-Seq长期以来一直是产生开放染色质全基因组图谱的金标准,上图中蓝色显示。 因为DNAse-Seq需要数千万个细胞,并且在技术上具有挑战性,所以它的实用性受到限制。 Active Motif的ATAC-Seq(上图中以绿色显示)仅使用50,000个细胞,并提供与DNAse-Seq相当的数据。
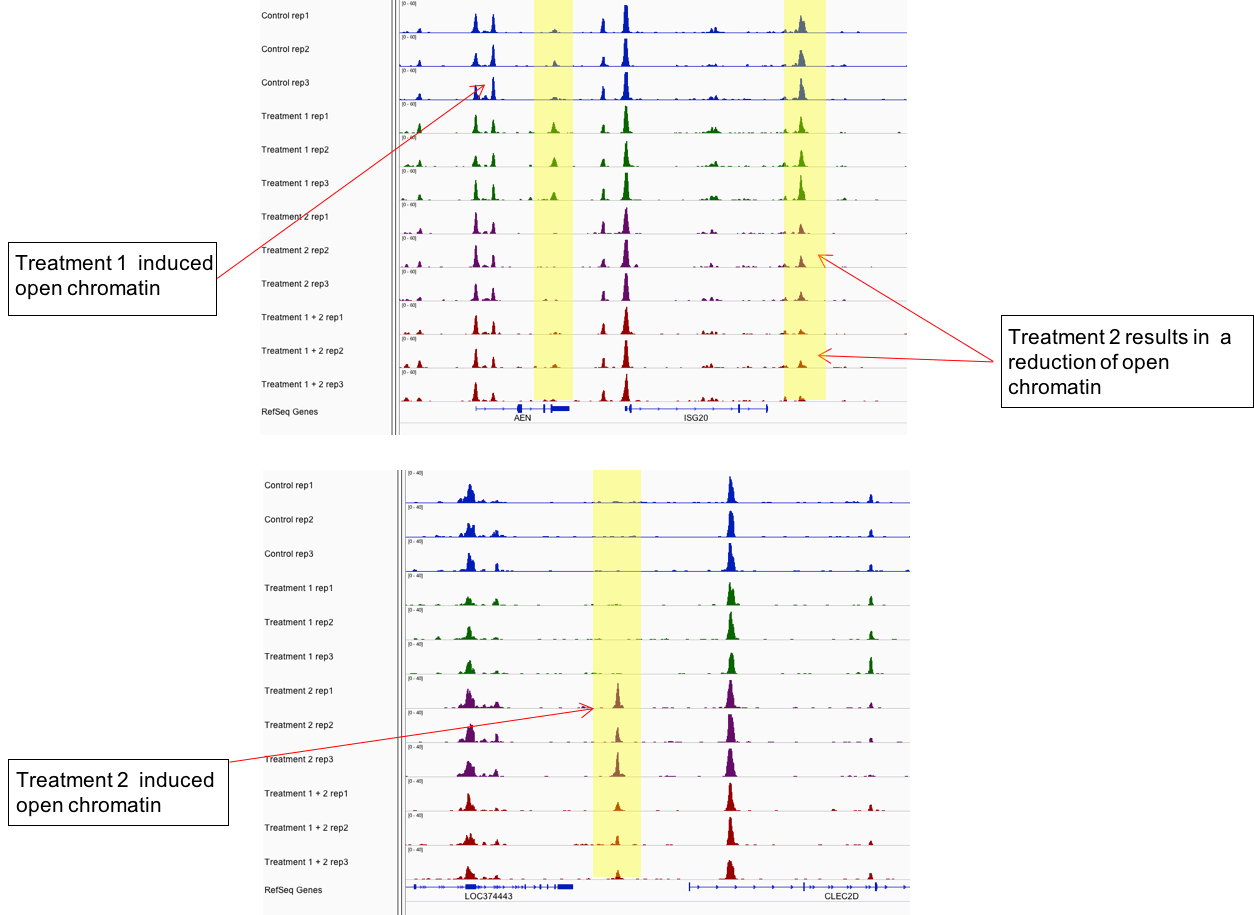
图2:Active Motif的ATAC-Seq分析通过识别差异开放的染色质区域来区分样品组。
上面的示例显示了来自4个不同样本的ATAC-Seq数据,每个样本3次重复。 差异开放区域以黄色突出显示。
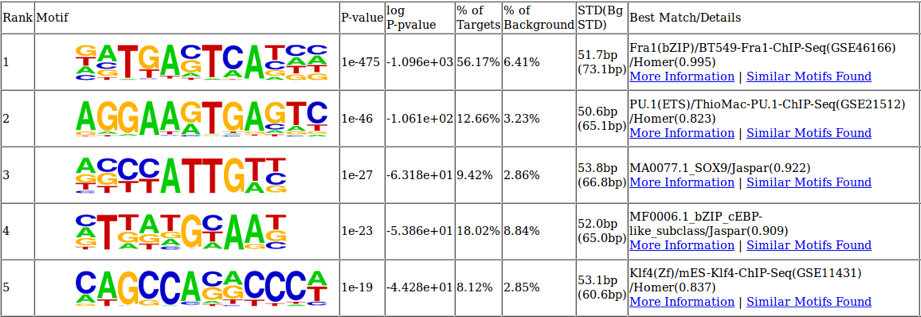
图3:使用ATAC-Seq识别重要的转录因子结合位点
可以分析差异开放的染色质区域的内在DNA序列,以鉴定最富集的转录因子结合位点。 在该细胞系统中,两个最丰富的结合基序也与B细胞生物学有关。 Fra1在B细胞激活后迅速上调,而PU.1是B细胞分化的关键调节因子。
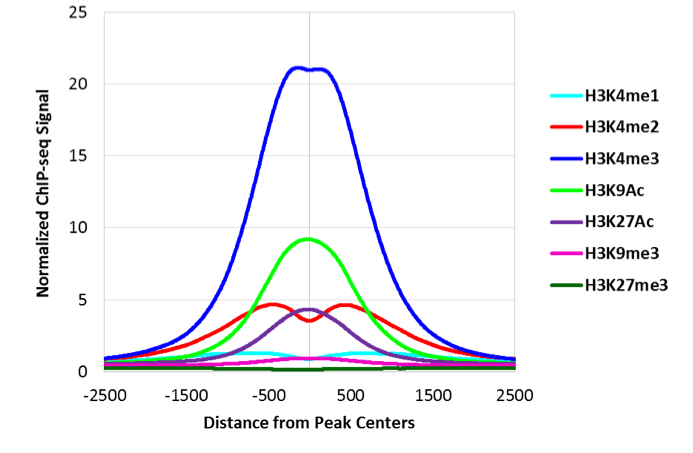
图4:注释的启动子上相对于ATAC-Seq峰的组蛋白修饰分布
将ATAC-Seq数据与不同的组蛋白修饰ChIP-Seq数据集进行比较发现,启动子处的ATAC-Seq峰富含最多的H3K4me3和H3K9Ac。
图5:带注释的启动子外,相对于ATAC-Seq峰的组蛋白修饰分布
启动子外的ATAC-Seq峰富含所有活跃标记,包括增强子标记H3K27Ac和H3K4me1。
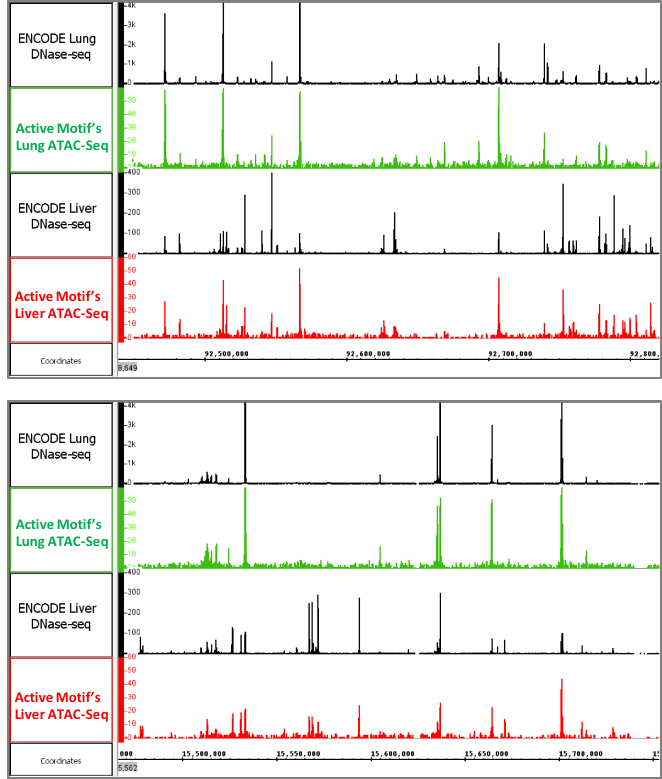
图6: Active Motif的组织样本ATAC-Seq数据
上图显示了使用冷冻的小鼠肝和肺组织的ATAC-Seq数据。 开放的染色质图谱类似于由ENCODE产生的DNAse-Seq图谱。
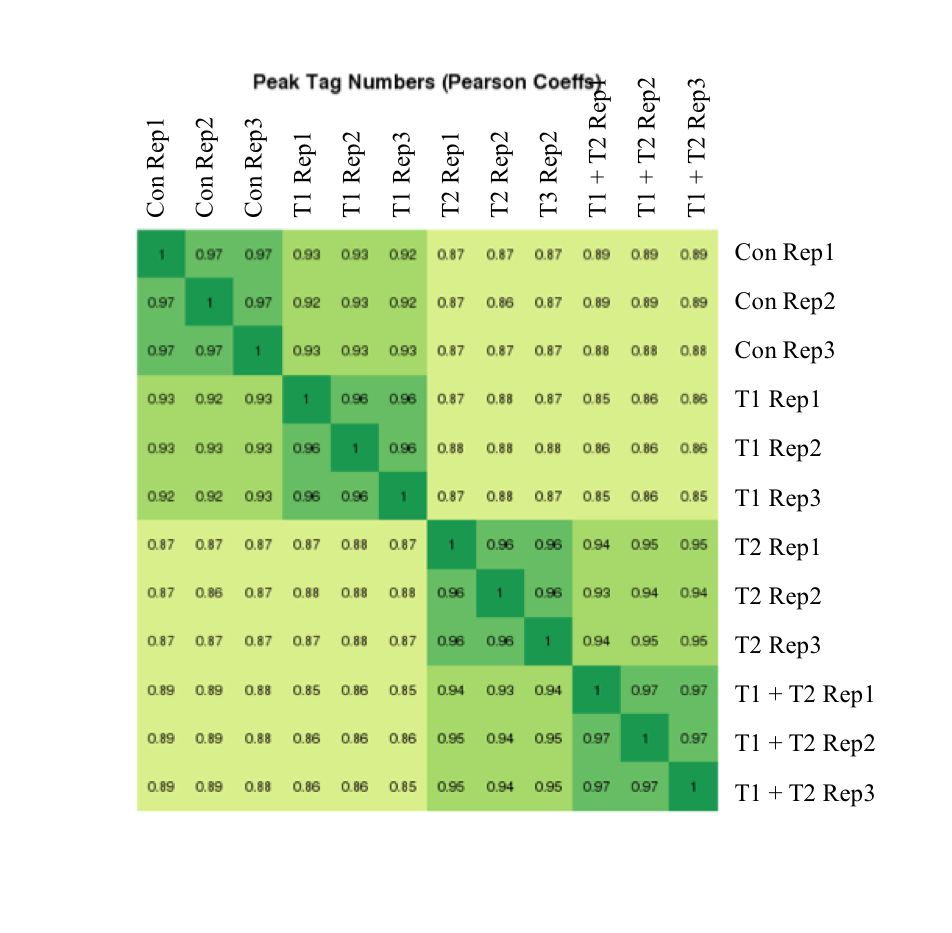
图7:Active Motif的ATAC-Seq数据显示出很高的重现性
使用未经处理或在三种不同条件下处理的细胞系进行上述实验,每种条件重复三次。 相关系数显示在热图中。 重复的相关系数至少为0.96。 热图显示,样品按预期分为四个不同的组。
ATAC-Seq Quality Measures
有多种方法可以评估ATAC-Seq数据的质量。最重要的两个是FRiP得分和峰数量。
- FRiP分数:全称Fraction of Reads in Peaks,代表的是与peaks有交叉的reads占总reads数的百分比。 它是对开放区域富集的一种度量,也可以看作是信噪比的度量,其中信号为比对到peak上的reads,噪音为比对到peak外的reads。 FRiP分数根据细胞类型而有所不同。 FRiP分数> 30%是成功的良好指标。 但是,对于一些难处理的样本,只要样本之间具有一致性,较低的FRiP分数也是可以接受的。
- 峰数量:在ATAC-Seq数据中识别出的峰的数量。 像ENCODE这样的数据库建议数据识别出50,000个以上的峰。 但是,这取决于细胞类型,组织和细胞健康状况。

图1:Active Motif优化的ATAC-Seq方法得到更高的FRiP分数使用标准ATAC-Seq方法*(蓝色),Omni ATAC-Seq方法#(橙色)或Active Motif的ATAC-Seq方法(紫色)处理人胚胎祖细胞(4D20.8细胞系和原代细胞),人原代内皮细胞,人间充质干细胞和大鼠原代心肌细胞。 使用Active Motif的ATAC-Seq方法处理的样品始终能在各种样品中鉴定出更多峰。

图2:Active Motif优化的ATAC-Seq实验方法使得峰的数量增加。使用标准ATAC-Seq方法*(蓝色),Omni ATAC-Seq方法#(橙色)或Active Motif的ATAC-Seq方法(紫色)处理人胚胎祖细胞(4D20.8细胞系和原代细胞),人原代内皮细胞,人间充质干细胞和大鼠原代心肌细胞。 使用Active Motif的ATAC-Seq方法处理的样品始终能在各种样品中鉴定出更多峰。
ATAC-Seq和Omni ATAC-Seq已发表的文献
- Buenrostro, J.D. et al. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position, Nature Methods. 2013; 10:1213-1218.
- Corces, M.R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues, Nature Methods. 2017; 14:959-962.